Statistieken buiten geoms
Gevorderde datavisualisatie met ggplot2

Rick Scavetta
Founder, Scavetta Academy
Basisplot
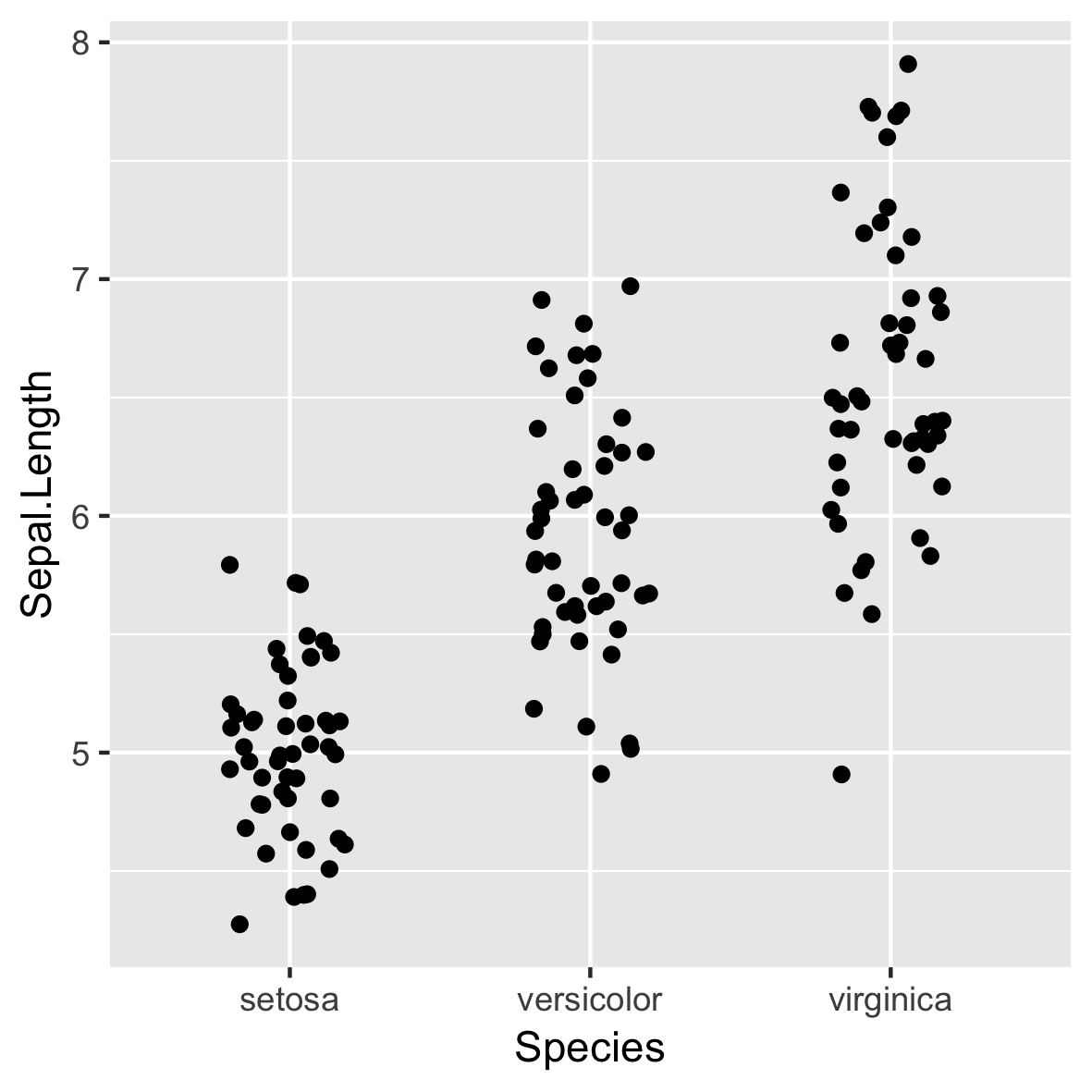
ggplot(iris, aes(x = Species,
y = Sepal.Length)) +
geom_jitter(width = 0.2)
Statistieken berekenen
set.seed(123)
xx <- rnorm(100)
mean(xx)
[1] 0.09040591
mean(xx) + (sd(xx) * c(-1, 1))
[1] -0.822410 1.003222
Statistieken berekenen
set.seed(123)
xx <- rnorm(100)
# Hmisc
library(Hmisc)
smean.sdl(xx, mult = 1)
Mean Lower Upper
0.09040591 -0.82240997 1.00322179
# ggplot2
mean_sdl(xx, mult = 1)
y ymin ymax
1 0.09040591 -0.82241 1.003222
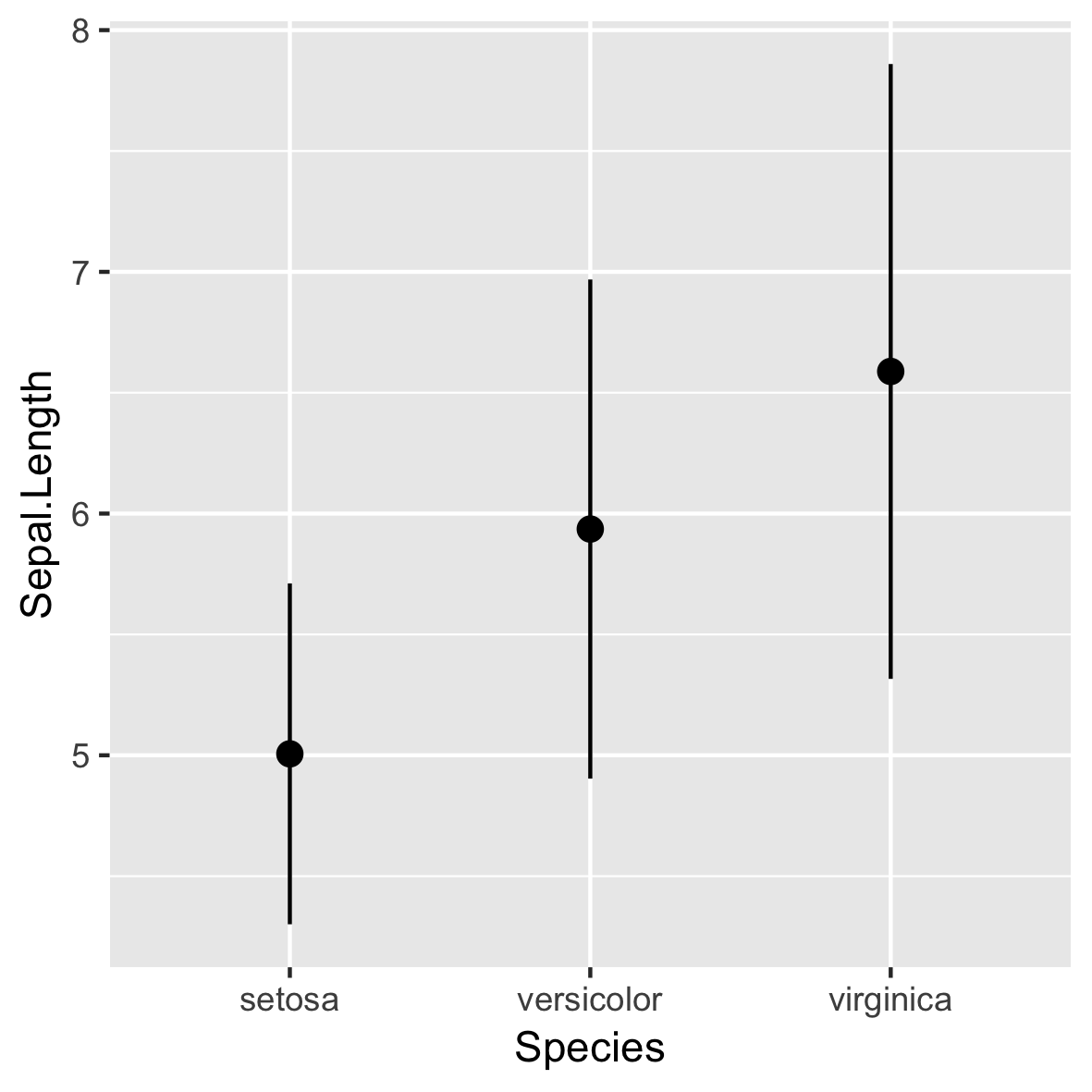
stat_summary()
ggplot(iris, aes(x = Species,
y = Sepal.Length)) +
stat_summary(fun.data = mean_sdl,
fun.args = list(mult = 1))
- Gebruikt standaard
geom_pointrange()
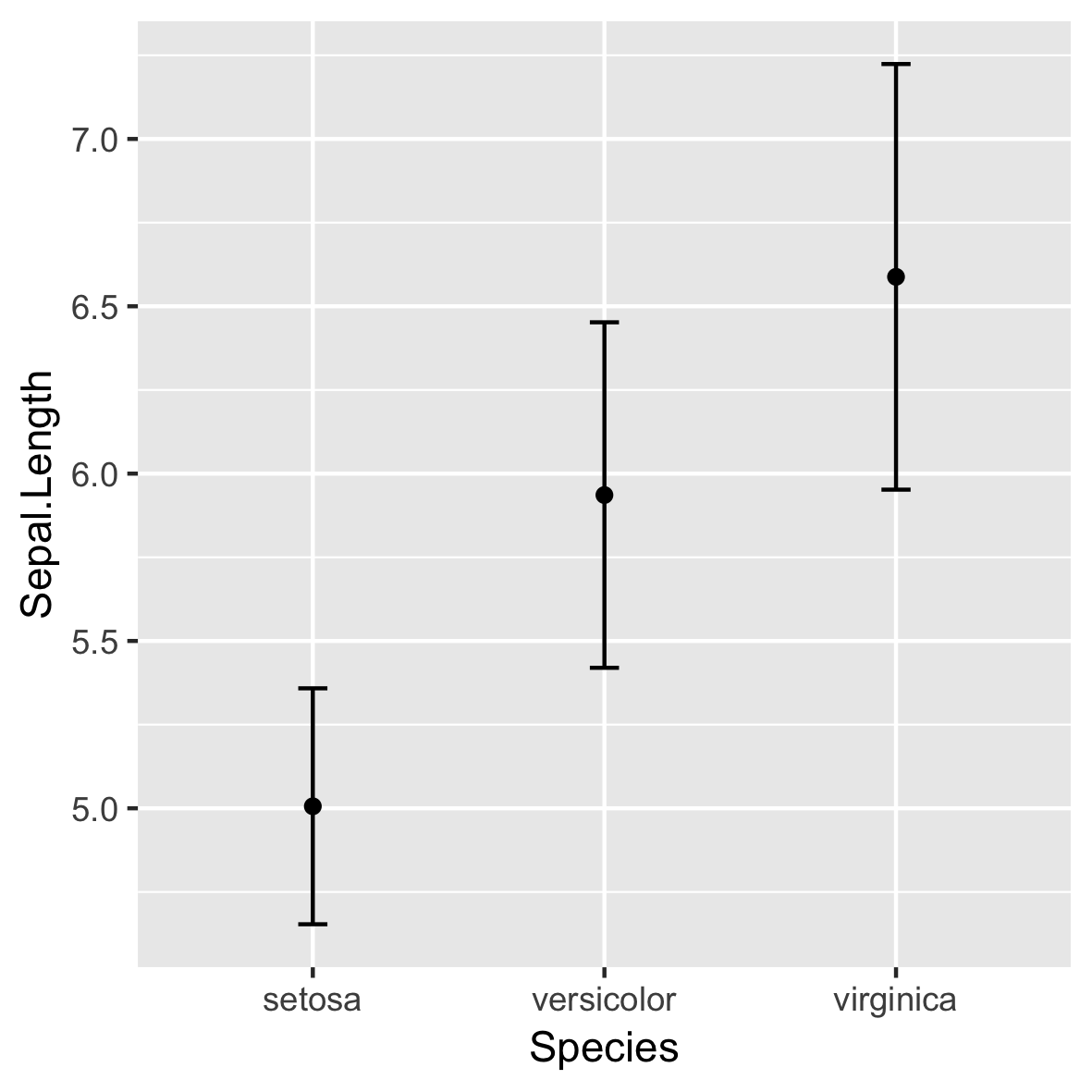
stat_summary()
ggplot(iris, aes(x = Species,
y = Sepal.Length)) +
stat_summary(fun = mean,
geom = "point") +
stat_summary(fun.data = mean_sdl,
fun.args = list(mult = 1),
geom = "errorbar",
width = 0.1)
Niet aanbevolen!
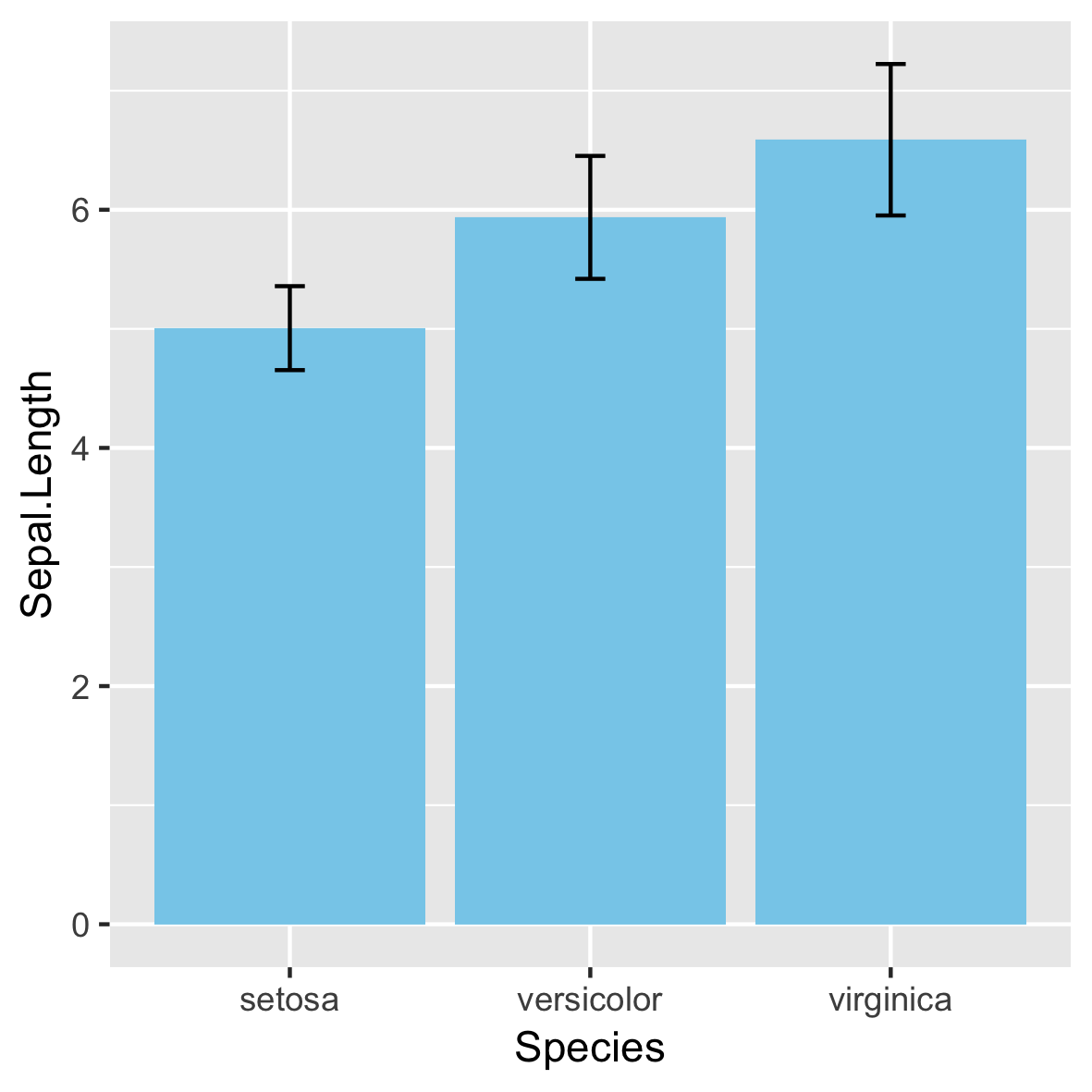
95%-betrouwbaarheidsinterval
ERR <- qt(0.975, length(xx) - 1) * (sd(xx) / sqrt(length(xx)))
mean(xx)
0.09040591
mean(xx) + (ERR * c(-1, 1)) # 95% CI
-0.09071657 0.27152838
mean_cl_normal(xx)
y ymin ymax
0.09040591 -0.09071657 0.2715284
Andere stat_-functies
stat_ |
Beschrijving |
|---|---|
stat_summary() |
vat y-waarden samen per unieke x. |
stat_function() |
berekent y uit een functie van x. |
stat_qq() |
berekent voor een quantile-quantile-plot. |
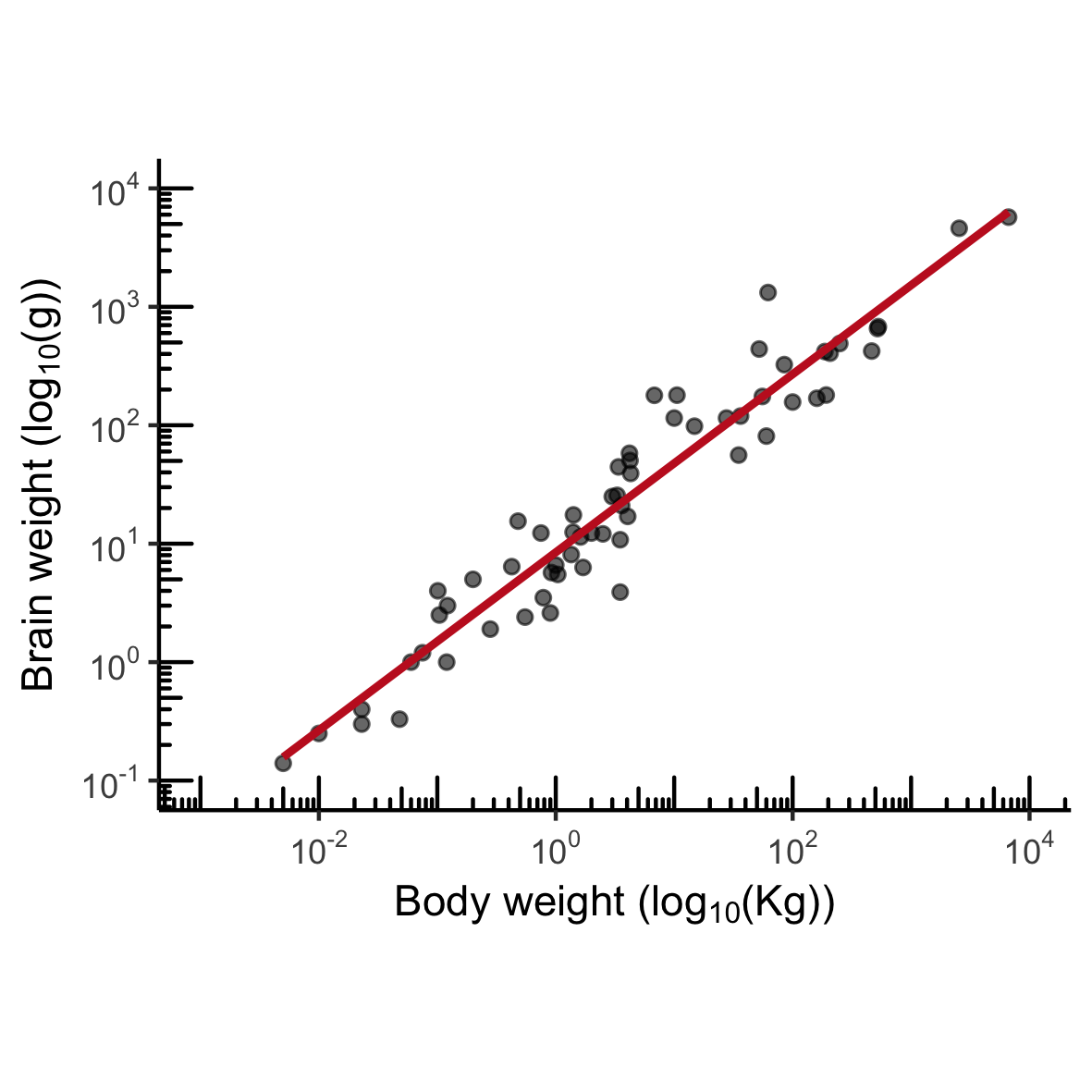
MASS::mammals
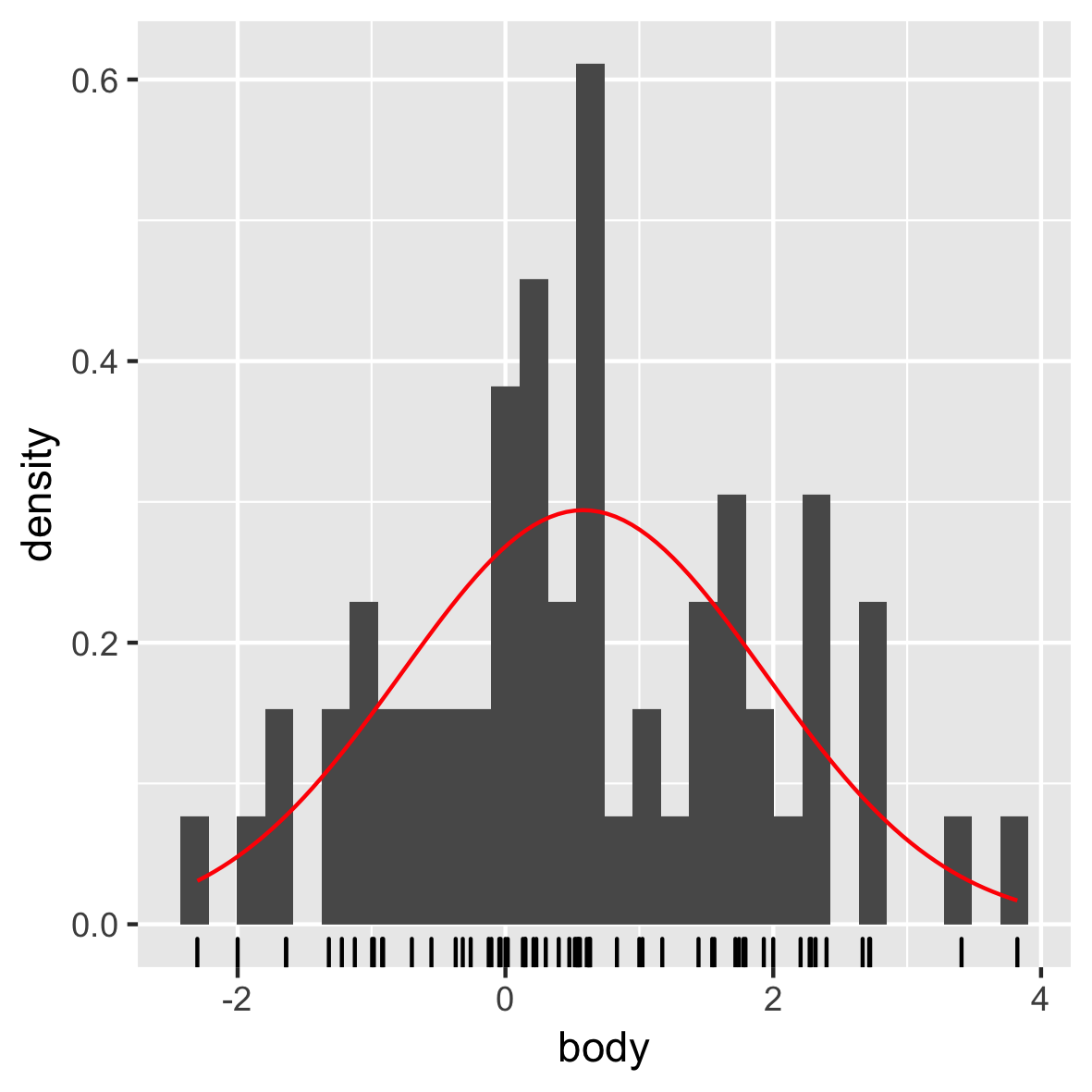
Normale verdeling
mam.new <- data.frame(body = log10(mammals$body))
ggplot(mam.new, aes(x = body)) +
geom_histogram(aes( y = ..density..)) +
geom_rug() +
stat_function(fun = dnorm, color = "red",
args = list(mean = mean(mam.new$body),
sd = sd(mam.new$body)))
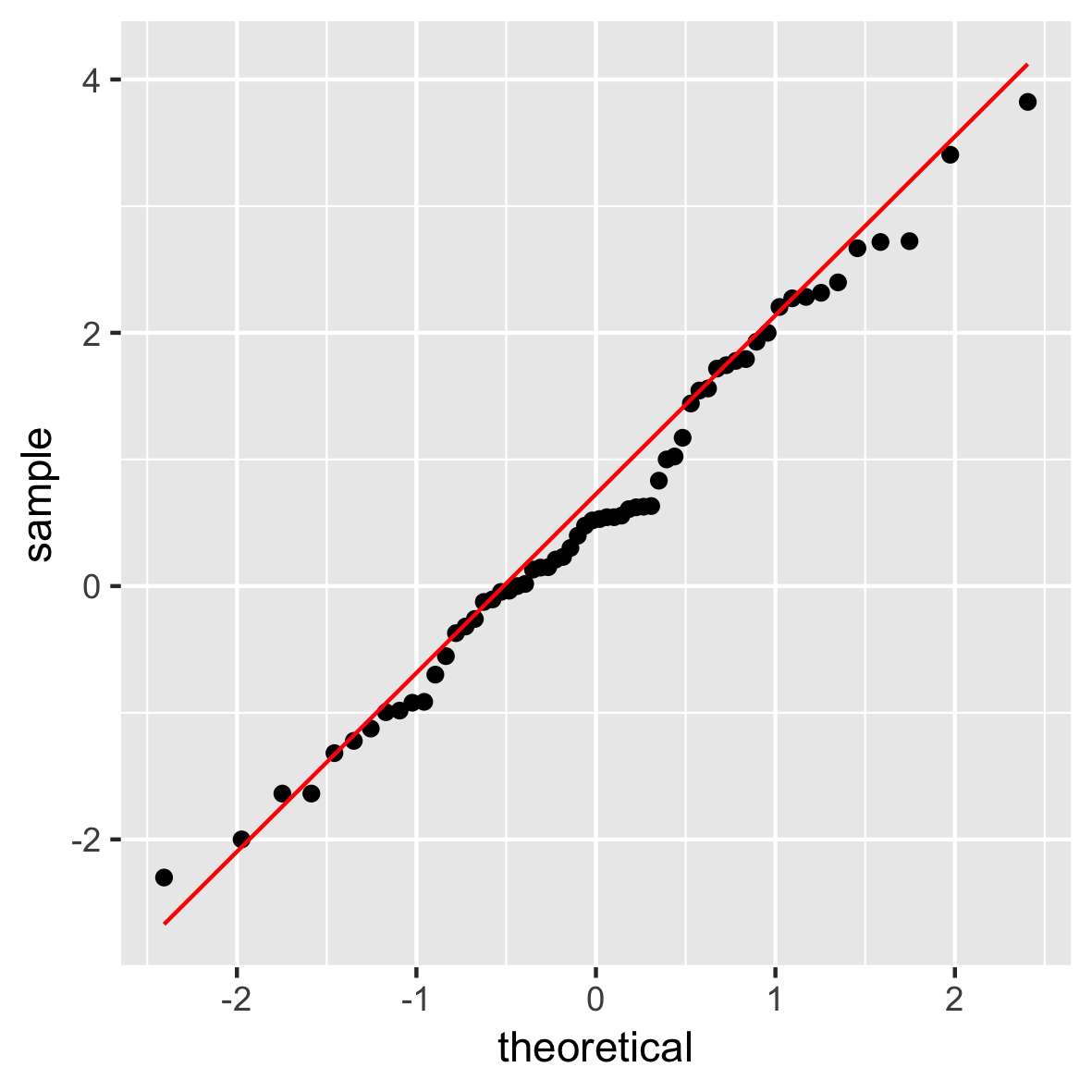
QQ-plot
ggplot(mam.new, aes(sample = body)) +
stat_qq() +
geom_qq_line(col = "red")
Jij bent aan de beurt!
Gevorderde datavisualisatie met ggplot2

