Gevoeligheidsanalyse
Monte Carlo-simulaties in Python

Izzy Weber
Curriculum Manager, DataCamp
Gevoeligheidsanalyse
Helpt de impact van de invoerbereiken te begrijpen
Laat patronen of trends zien in tabellen of plots
Als we de waarden voor bmi en hdl verhogen of verlagen met een Monte Carlo-simulatie, hoe veranderen dan de voorspelde y-waarden (ziekteprogressie)?
Parameters definiëren
cov_dia = dia[["age", "bmi", "bp", "tc", "ldl", "hdl", "tch", "ltg", "glu"]].cov()
mean_dia = dia[["age", "bmi", "bp", "tc", "ldl", "hdl", "tch", "ltg", "glu"]].mean()
De simulatiefunctie definiëren
def simulate_bmi_hdl(cov_dia, mean_list):list_ys = [] for i in range(50): simulation_results = st.multivariate_normal.rvs(mean=mean_list, size=500, cov=cov_dia) df_results = pd.DataFrame(simulation_results, columns=["age","bmi","bp","tc","ldl","hdl","tch","ltg","glu"]) predicted_y = regr_model.predict(df_results) df_y = pd.DataFrame(predicted_y, columns=["predicted_y"]) df_summary = pd.concat([df_results, df_y], axis=1) y = np.mean(df_summary["predicted_y"]) list_ys.append(y)return(np.mean(list_ys))
Simulaties uitvoeren met een bereik aan invoerparameters
hdl = [] bmi = [] simu_y = [] for mean_hdl_inc in np.arange(-20, 50, 30): for mean_bmi_inc in np.arange(-7, 11, 3):mean_list = mean_dia + np.array([0, mean_bmi_inc, 0, 0, 0, mean_hdl_inc, 0, 0, 0]) hdl.append(mean_hdl_inc) bmi.append(mean_bmi_inc)mean_y = simulate_bmi_hdl(cov_dia, mean_list)simu_y.append(mean_y)df_sa = pd.concat([pd.Series(hdl), pd.Series(bmi), pd.Series(simu_y)], axis=1)df_sa.columns = ["hdl_inc", "bmi_inc", "y"]
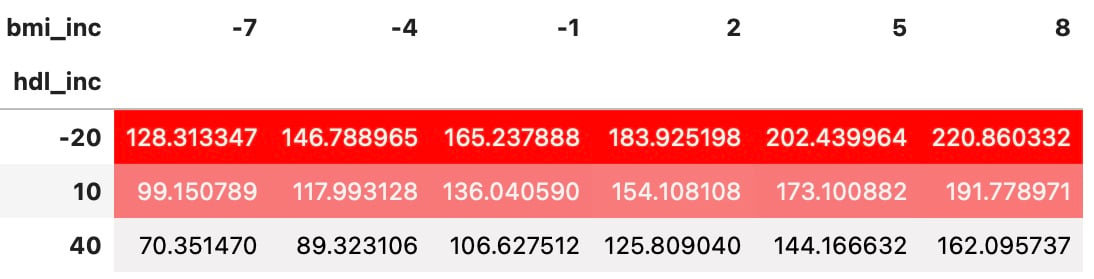
Gestylede DataFrames van resultaten gevoeligheidsanalyse
df_sa.sort_values(by=['hdl_inc', 'bmi_inc']).pivot(index='hdl_inc',
columns='bmi_inc',
values='y').style.background_gradient(
cmap=sns.light_palette("red", as_cmap=True))
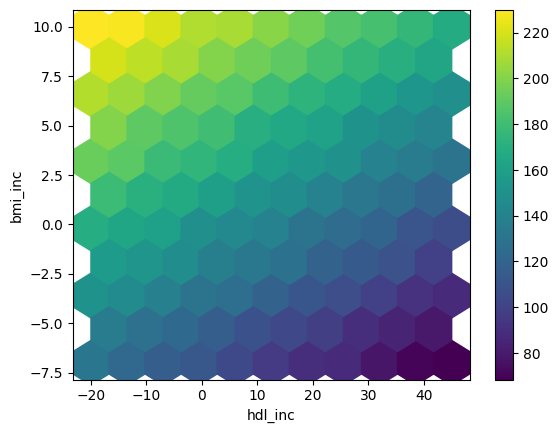
Hexbin-plot voor resultaten van gevoeligheidsanalyse
df_sa.plot.hexbin(x='hdl_inc',y='bmi_inc', C='y',
reduce_C_function=np.mean,
gridsize=10, cmap="viridis",
sharex=False)

Hexbin-plot voor een dense parameterspace
Laten we oefenen!
Monte Carlo-simulaties in Python

