Analisi di sensitività
Simulazioni Monte Carlo in Python

Izzy Weber
Curriculum Manager, DataCamp
Analisi di sensitività
Aiuta a capire l’impatto dell’intervallo degli input
Mostra pattern o trend quando riassunti in tabelle o grafici
Se aumentiamo o diminuiamo i valori di bmi e hdl con una simulazione Monte Carlo, come cambiano i valori previsti di y (progressione della malattia)?
Definizione dei parametri
cov_dia = dia[["age", "bmi", "bp", "tc", "ldl", "hdl", "tch", "ltg", "glu"]].cov()
mean_dia = dia[["age", "bmi", "bp", "tc", "ldl", "hdl", "tch", "ltg", "glu"]].mean()
Definizione della funzione di simulazione
def simulate_bmi_hdl(cov_dia, mean_list):list_ys = [] for i in range(50): simulation_results = st.multivariate_normal.rvs(mean=mean_list, size=500, cov=cov_dia) df_results = pd.DataFrame(simulation_results, columns=["age","bmi","bp","tc","ldl","hdl","tch","ltg","glu"]) predicted_y = regr_model.predict(df_results) df_y = pd.DataFrame(predicted_y, columns=["predicted_y"]) df_summary = pd.concat([df_results, df_y], axis=1) y = np.mean(df_summary["predicted_y"]) list_ys.append(y)return(np.mean(list_ys))
Simula con un intervallo di parametri in input
hdl = [] bmi = [] simu_y = [] for mean_hdl_inc in np.arange(-20, 50, 30): for mean_bmi_inc in np.arange(-7, 11, 3):mean_list = mean_dia + np.array([0, mean_bmi_inc, 0, 0, 0, mean_hdl_inc, 0, 0, 0]) hdl.append(mean_hdl_inc) bmi.append(mean_bmi_inc)mean_y = simulate_bmi_hdl(cov_dia, mean_list)simu_y.append(mean_y)df_sa = pd.concat([pd.Series(hdl), pd.Series(bmi), pd.Series(simu_y)], axis=1)df_sa.columns = ["hdl_inc", "bmi_inc", "y"]
DataFrame stilizzati dei risultati dell’analisi di sensitività
df_sa.sort_values(by=['hdl_inc', 'bmi_inc']).pivot(index='hdl_inc',
columns='bmi_inc',
values='y').style.background_gradient(
cmap=sns.light_palette("red", as_cmap=True))
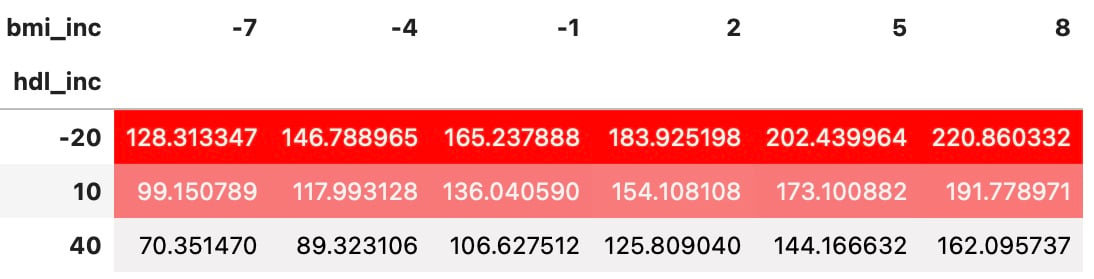
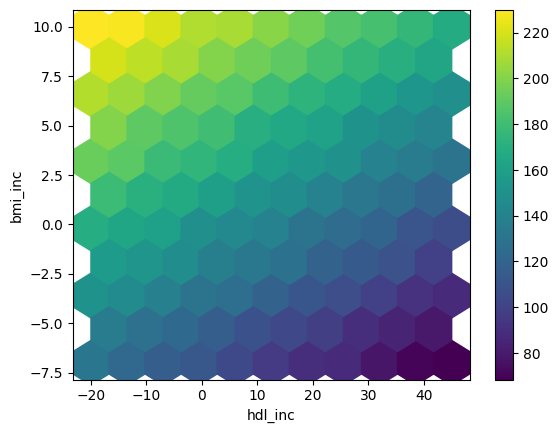
Grafico hexbin per i risultati dell’analisi di sensitività
df_sa.plot.hexbin(x='hdl_inc',y='bmi_inc', C='y',
reduce_C_function=np.mean,
gridsize=10, cmap="viridis",
sharex=False)

Grafico hexbin per spazio dei parametri denso
Ayo berlatih!
Simulazioni Monte Carlo in Python

