Replicare la variabilità dei dati
Gestione dei dati mancanti con imputazioni in R

Michal Oleszak
Machine Learning Engineer
Variabilità nei dati imputati
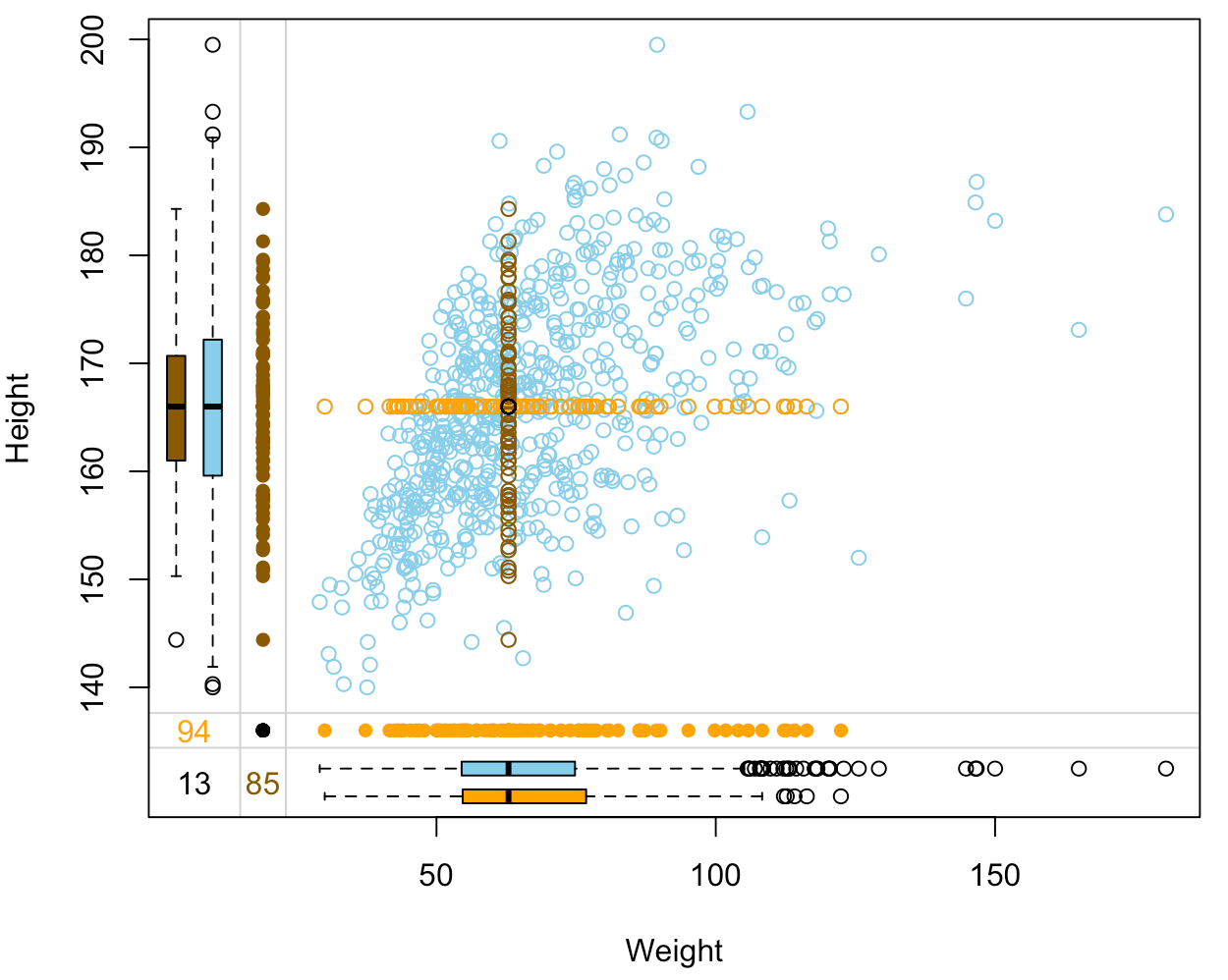
- Nessuna variabilità nei dati imputati.
- Vogliamo che l'imputazione replichi la variabilità dei dati osservati.
- Nell'imputazione basata su modelli, gli stessi predittori danno lo stesso valore imputato.
- Soluzione: campionare da distribuzioni condizionali.
Cos'è una previsione
La maggior parte dei modelli statistici stima la distribuzione condizionale della variabile risposta:
$p(y|X)$
Per fare una singola previsione, si riassume la distribuzione condizionale:
- Regressione lineare: valore atteso della distribuzione condizionale.
- Regressione logistica: classe con probabilità più alta.
Invece, possiamo campionare da queste distribuzioni per aumentare la variabilità.
Campionare da distribuzioni condizionali
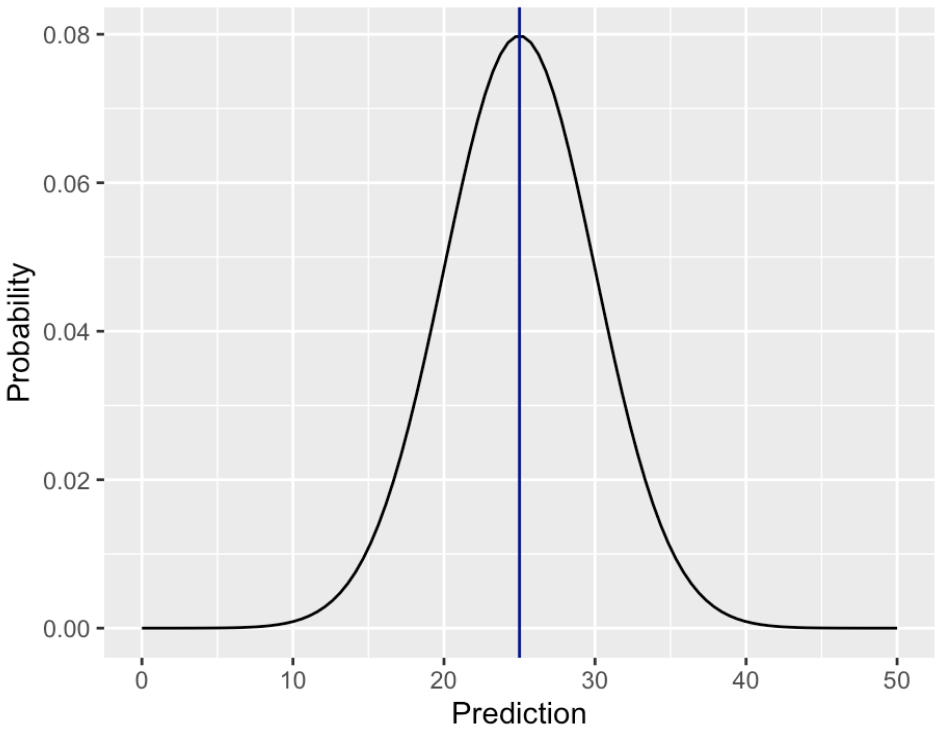
Campionare da distribuzioni condizionali
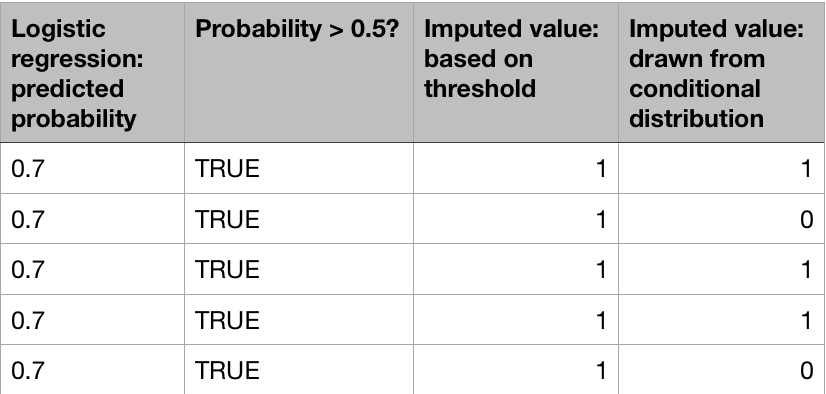
Imputazione con regressione logistica
Compito: imputa PhysActive dai dati nhanes con regressione logistica.
nhanes_imp <- hotdeck(nhanes)
missing_physactive <- is.na(nhanes$PhysActive)
Imputazione con regressione logistica
Compito: imputa PhysActive dai dati nhanes con regressione logistica.
nhanes_imp <- hotdeck(nhanes)
missing_physactive <- is.na(nhanes$PhysActive)
logreg_model <- glm(PhysActive ~ Age + Weight + Pulse,
data = nhanes_imp, family = binomial)
Imputazione con regressione logistica
Compito: imputa PhysActive dai dati nhanes con regressione logistica.
nhanes_imp <- hotdeck(nhanes)
missing_physactive <- is.na(nhanes$PhysActive)
logreg_model <- glm(PhysActive ~ Age + Weight + Pulse,
data = nhanes_imp, family = binomial)
preds <- predict(logreg_model, type = "response")
Imputazione con regressione logistica
Compito: imputa PhysActive dai dati nhanes con regressione logistica.
nhanes_imp <- hotdeck(nhanes)
missing_physactive <- is.na(nhanes$PhysActive)
logreg_model <- glm(PhysActive ~ Age + Weight + Pulse,
data = nhanes_imp, family = binomial)
preds <- predict(logreg_model, type = "response")
preds <- ifelse(preds >= 0.5, 1, 0)
Imputazione con regressione logistica
Compito: imputa PhysActive dai dati nhanes con regressione logistica.
nhanes_imp <- hotdeck(nhanes)
missing_physactive <- is.na(nhanes$PhysActive)
logreg_model <- glm(PhysActive ~ Age + Weight + Pulse,
data = nhanes_imp, family = binomial)
preds <- predict(logreg_model, type = "response")
preds <- ifelse(preds >= 0.5, 1, 0)
nhanes_imp[missing_physactive, "PhysActive"] <- preds[missing_physactive]
Imputazione con regressione logistica
Variabilità dei dati imputati:
table(preds[missing_physactive])
1
26
Variabilità dei dati PhysActive osservati:
table(nhanes$PhysActive)
0 1
181 610
Campionare dalle probabilità di classe
nhanes_imp <- hotdeck(nhanes)
missing_physactive <- is.na(nhanes$PhysActive)
logreg_model <- glm(PhysActive ~ Age + Weight + Pulse,
data = nhanes_imp, family = binomial)
preds <- predict(logreg_model, type = "response")
preds <- ifelse(preds >= 0.5, 1, 0)
nhanes_imp[missing_physactive, "PhysActive"] <- preds[missing_physactive]
Campionare dalle probabilità di classe
nhanes_imp <- hotdeck(nhanes)
missing_physactive <- is.na(nhanes$PhysActive)
logreg_model <- glm(PhysActive ~ Age + Weight + Pulse,
data = nhanes_imp, family = binomial)
preds <- predict(logreg_model, type = "response")
nhanes_imp[missing_physactive, "PhysActive"] <- preds[missing_physactive]
Campionare dalle probabilità di classe
nhanes_imp <- hotdeck(nhanes)
missing_physactive <- is.na(nhanes$PhysActive)
logreg_model <- glm(PhysActive ~ Age + Weight + Pulse,
data = nhanes_imp, family = binomial)
preds <- predict(logreg_model, type = "response")
preds <- rbinom(length(preds), size = 1, prob = preds)
nhanes_imp[missing_physactive, "PhysActive"] <- preds[missing_physactive]
Campionare dalle probabilità di classe
Variabilità dei dati imputati:
table(preds[missing_physactive])
0 1
5 21
Variabilità dei dati PhysActive osservati:
table(nhanes$PhysActive)
0 1
181 610
Ayo berlatih!
Gestione dei dati mancanti con imputazioni in R

