Assessing inference from imputed data in a modelling context
Dealing With Missing Data in R

Nicholas Tierney
Statistician
Exploring parameters of one model
lm(Temp ~ Ozone + Solar.R + Wind + Month + day, data = airquality)
Complete case analysis
Imputation using the imputed data from the last lesson
Combining the datasets together
#1. Complete cases aq_cc <- airquality %>% na.omit() %>% bind_shadow() %>% add_label_shadow()#2. Imputation using the imputed data from the last lesson aq_imp_lm <- bind_shadow(airquality) %>% add_label_shadow() %>% impute_lm(Ozone ~ Temp + Wind + Month + Day) %>% impute_lm(Solar.R ~ Temp + Wind + Month + Day)# 3. Bind the models together bound_models <- bind_rows(cc = aq_cc, imp_lm = aq_imp_lm, .id = "imp_model")
Combining the datasets together
bound_models
imp_model Ozone Solar.R Wind Temp Month Day Ozone_NA Solar.R_NA any_missing
cc 41 190 7.4 67 5 1 !NA !NA Not Missing
cc 36 118 8.0 72 5 2 !NA !NA Not Missing
cc 12 149 12.6 74 5 3 !NA !NA Not Missing
cc 18 313 11.5 62 5 4 !NA !NA Not Missing
cc 23 299 8.6 65 5 7 !NA !NA Not Missing
...
imp_lm 30 193 6.9 70 9 26 !NA !NA Not Missing
imp_lm NA 145 13.2 77 9 27 NA !NA Missing
imp_lm 14 191 14.3 75 9 28 !NA !NA Not Missing
imp_lm 18 131 8.0 76 9 29 !NA !NA Not Missing
imp_lm 20 223 11.5 68 9 30 !NA !NA Not Missing
Exploring the models
model_summary <- bound_models %>%
group_by(imp_model) %>%
nest() %>%
mutate(mod = map(data,
~lm(Temp ~ Ozone + Solar.R + Wind + Temp + Days + Month
data = .)),
res = map(mod, residuals),
pred = map(mod, predict),
tidy = map(mod, broom::tidy))
model_summary
# A tibble: 2 x 6
imp_model data mod res pred tidy
<chr> <list> <list> <list> <list> <list>
1 cc <tibble [111 × 13]> <S3: lm> <dbl [111]> <dbl [111]> <tibble [3 × 5]>
2 imp_lm <tibble [153 × 13]> <S3: lm> <dbl [153]> <dbl [153]> <tibble [3 × 5]>
Exploring coefficients of multiple models
model_summary %>%
select(imp_model,
tidy) %>%
unnest()
# A tibble: 6 x 6
imp_model term estimate std.error statistic p.value
<chr> <chr> <dbl> <dbl> <dbl> <dbl>
1 cc (Intercept) 68.5 1.53 44.8 1.31e-71
2 cc Ozone 0.194 0.0210 9.26 2.22e-15
3 cc Solar.R 0.00604 0.00766 0.789 4.32e- 1
4 imp_lm (Intercept) 67.2 1.30 51.5 2.68e-97
5 imp_lm Ozone 0.215 0.0180 12.0 1.40e-23
6 imp_lm Solar.R 0.00787 0.00630 1.25 2.13e- 1
Exploring residuals of multiple models
model_summary %>%
select(imp_model,
res) %>%
unnest() %>%
ggplot(aes(x = res,
fill = imp_model)) +
geom_histogram(position = "dodge")
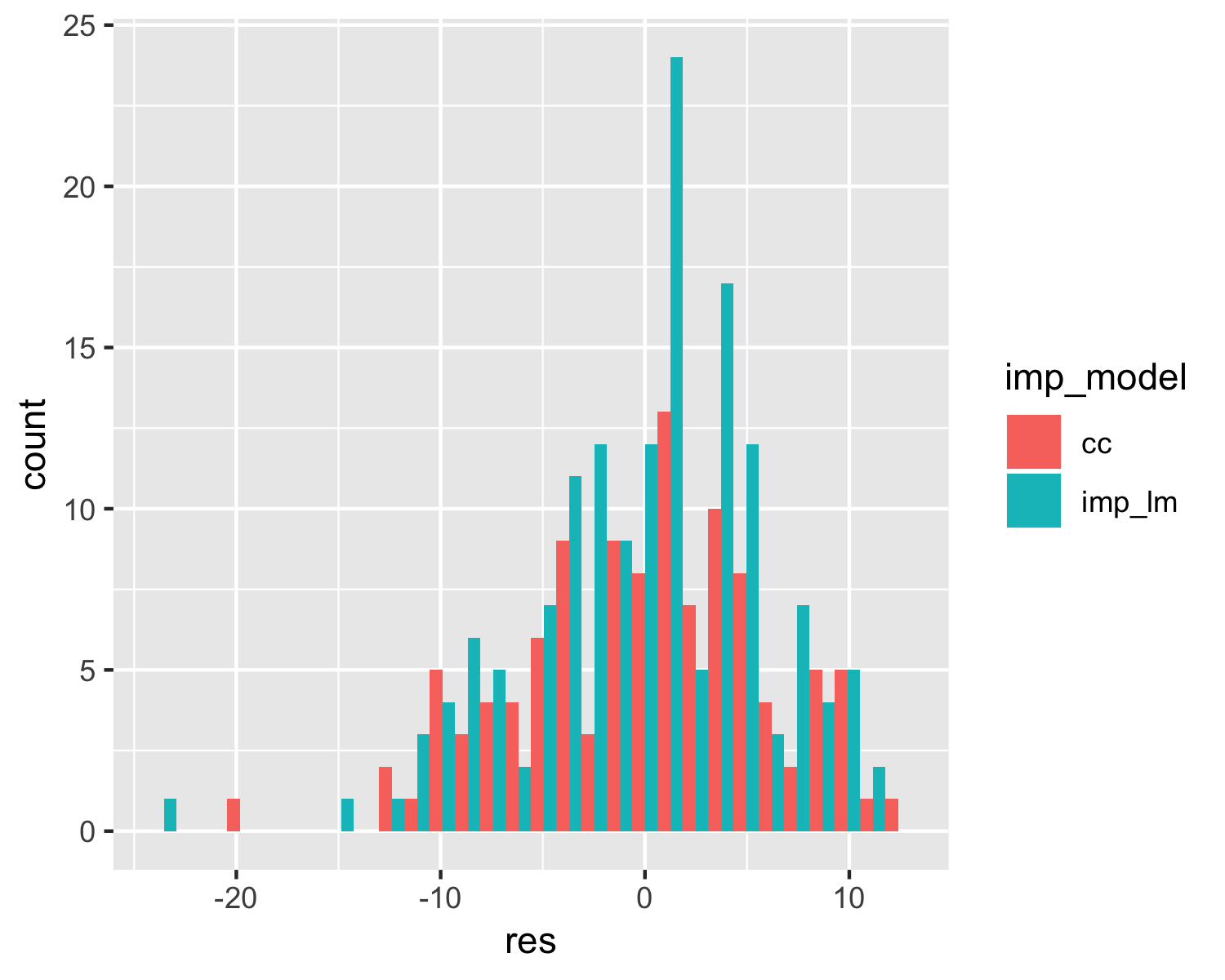
Exploring predictions of multiple models
model_summary %>%
select(imp_model,
pred) %>%
unnest() %>%
ggplot(aes(x = pred,
fill = imp_model)) +
geom_histogram(position = "dodge")
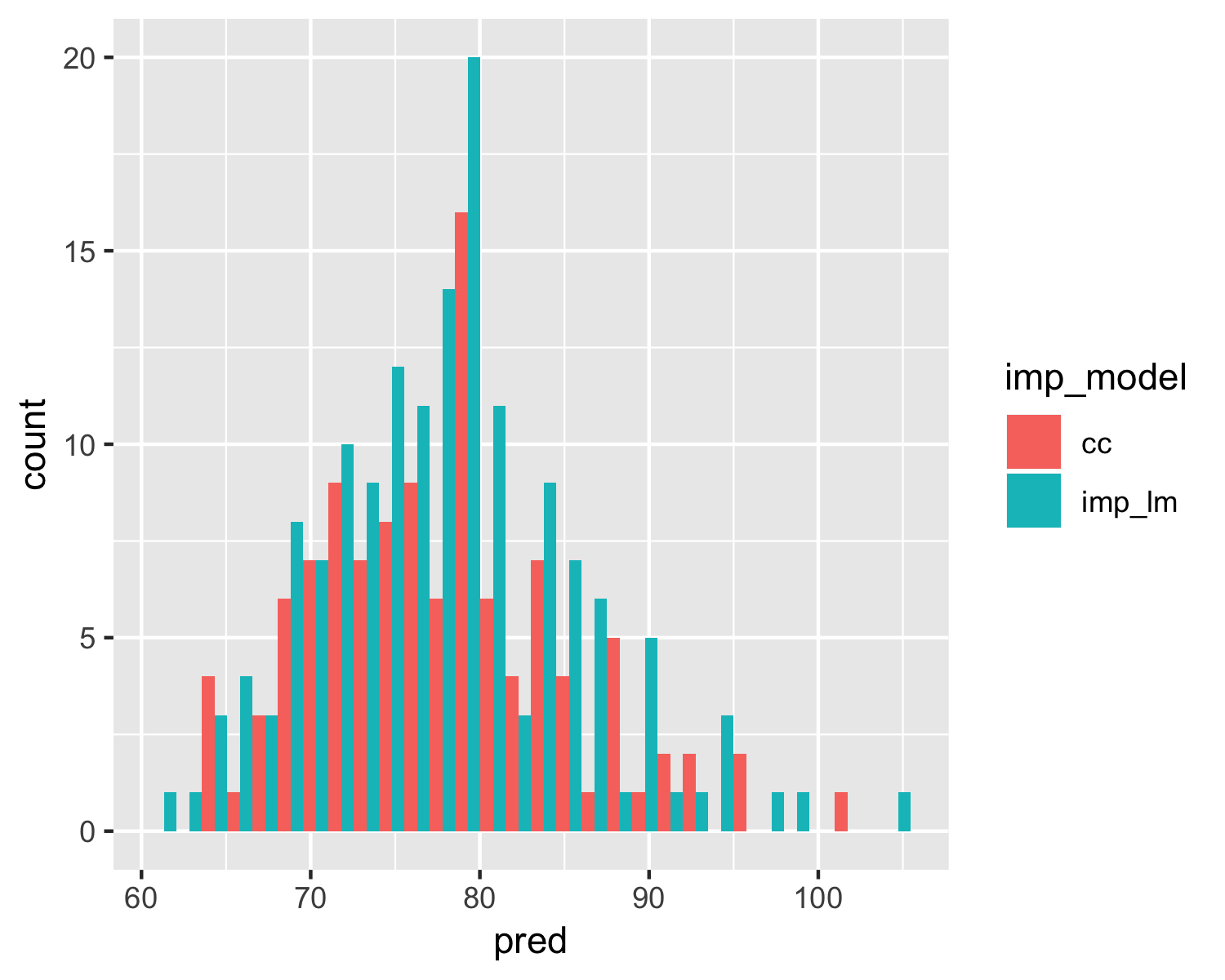
Let's practice!
Dealing With Missing Data in R

