Measuring cross-validation performance
Machine Learning in the Tidyverse

Dmitriy (Dima) Gorenshteyn
Lead Data Scientist, Memorial Sloan Kettering Cancer Center
Measuring Performance
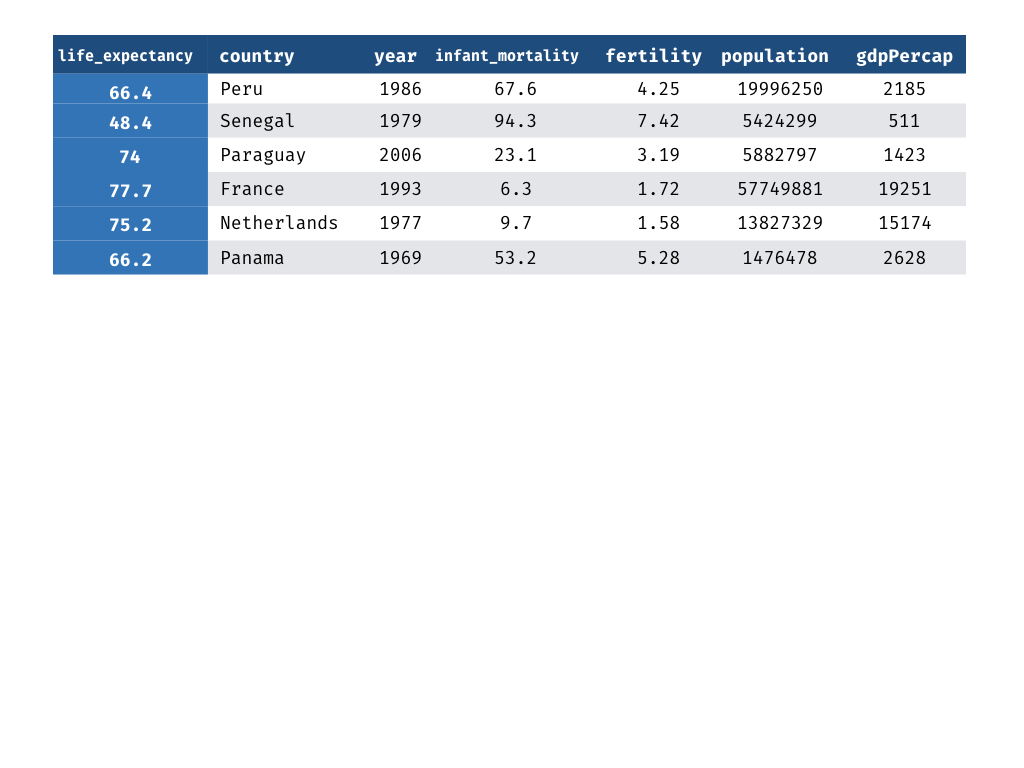
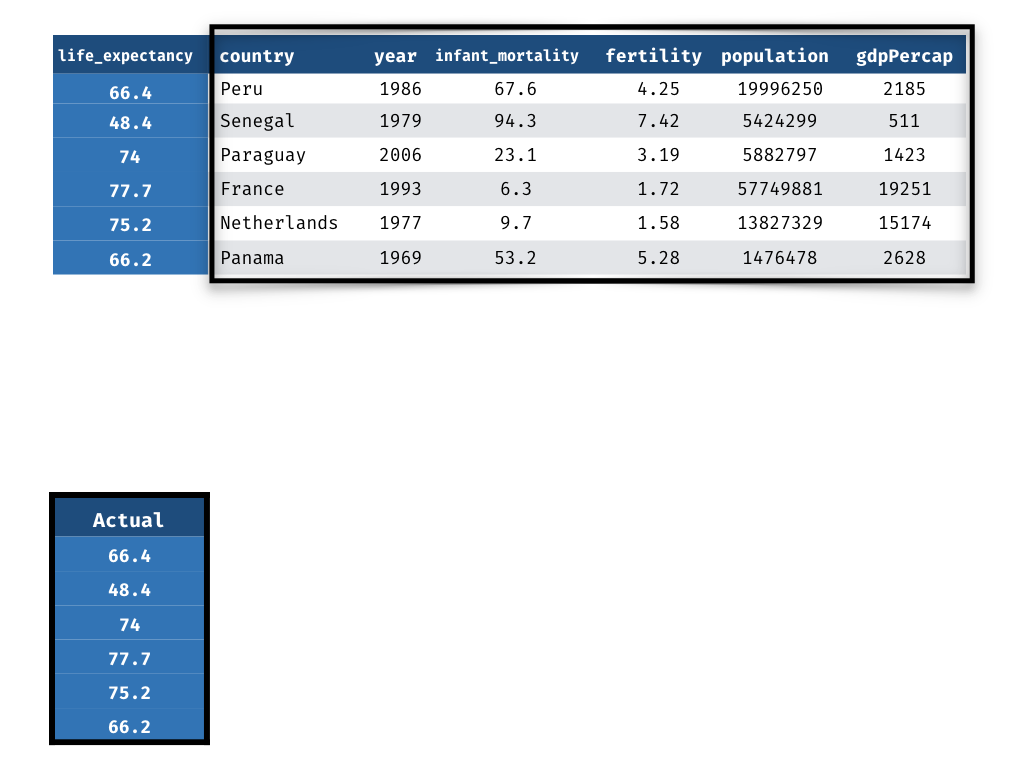
Measuring Performance - Truth
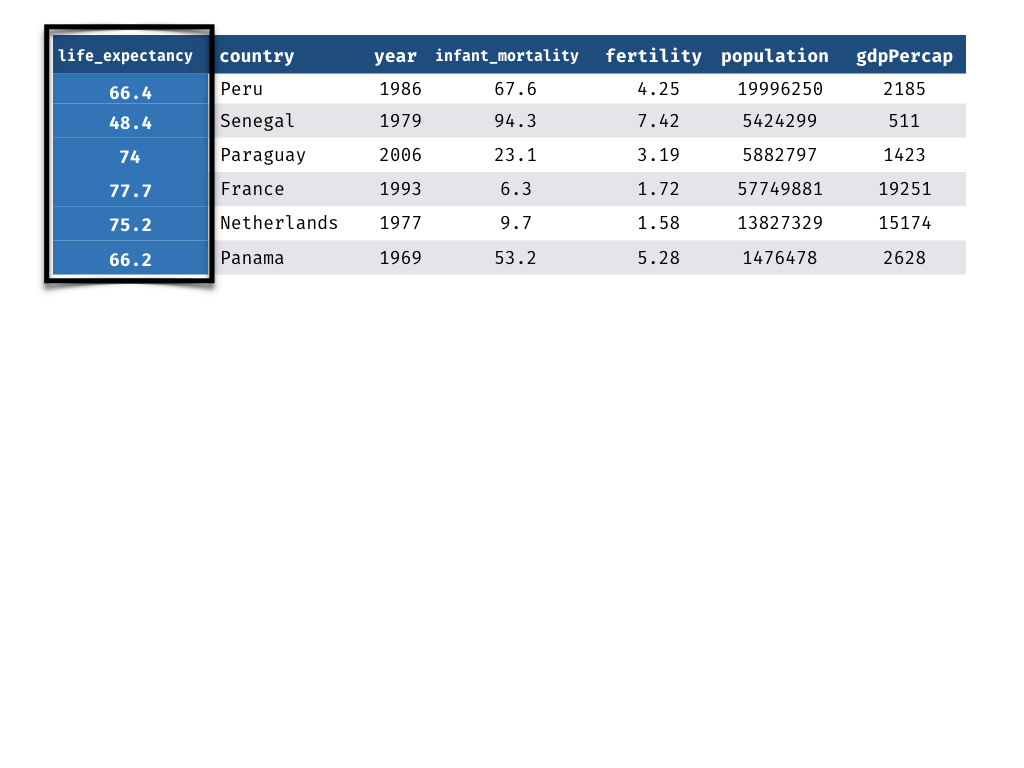
Measuring Performance - Truth
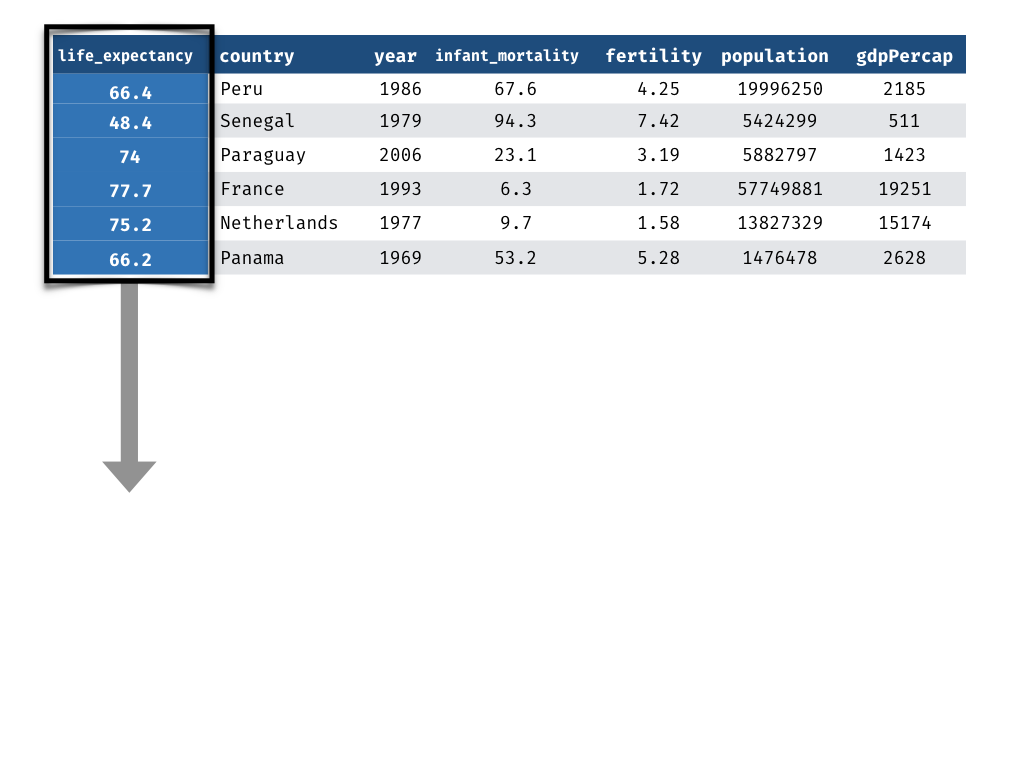
Measuring Performance - Truth
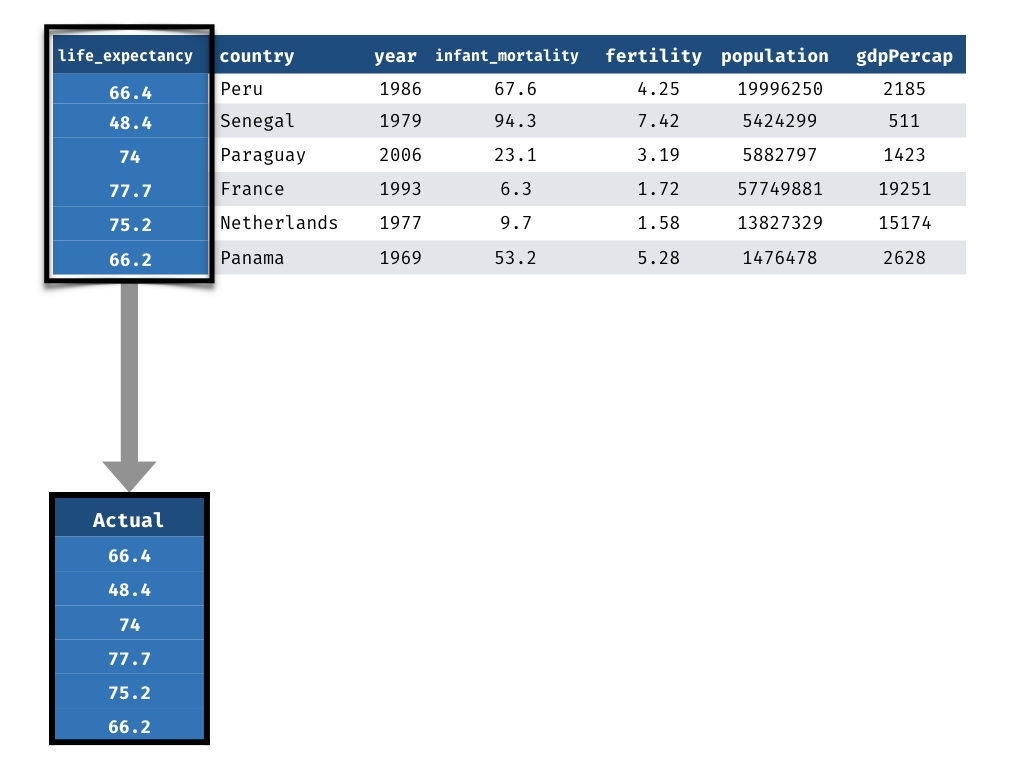
Measuring Performance - Prediction
Measuring Performance - Prediction
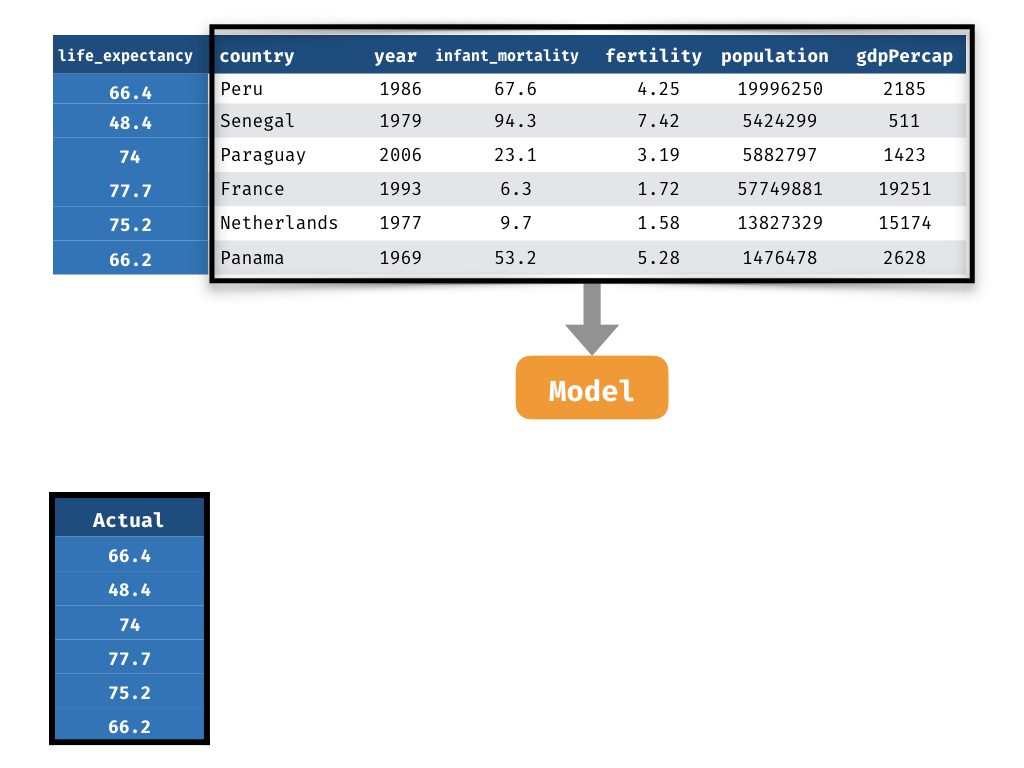
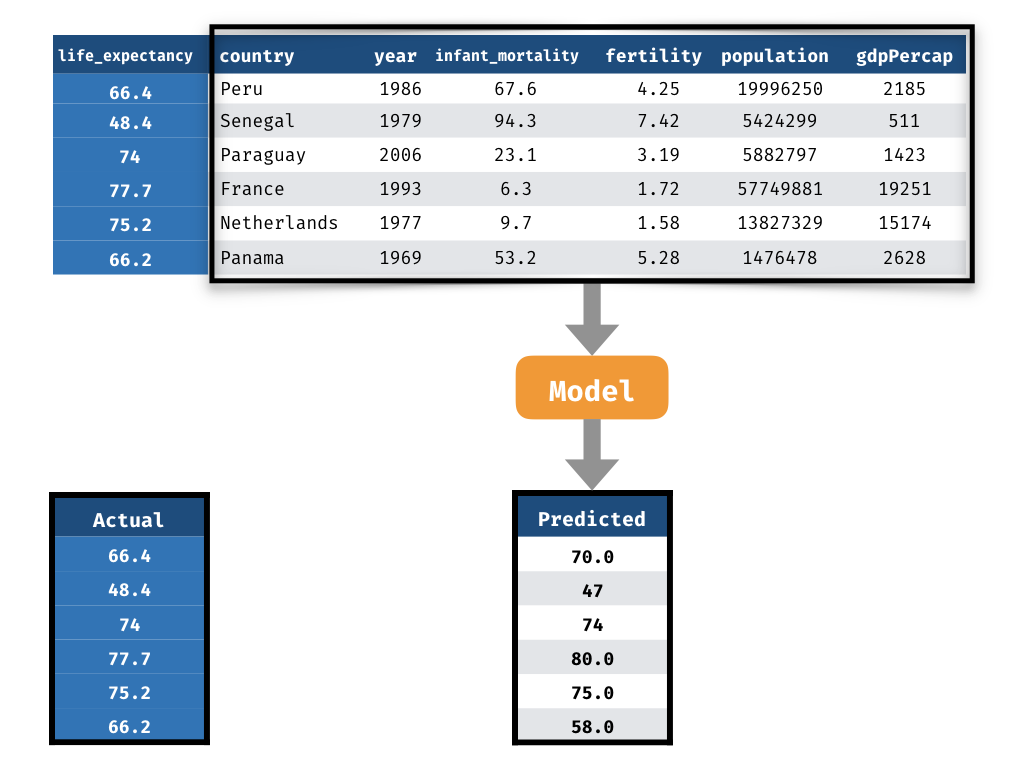
Measuring Performance - Prediction

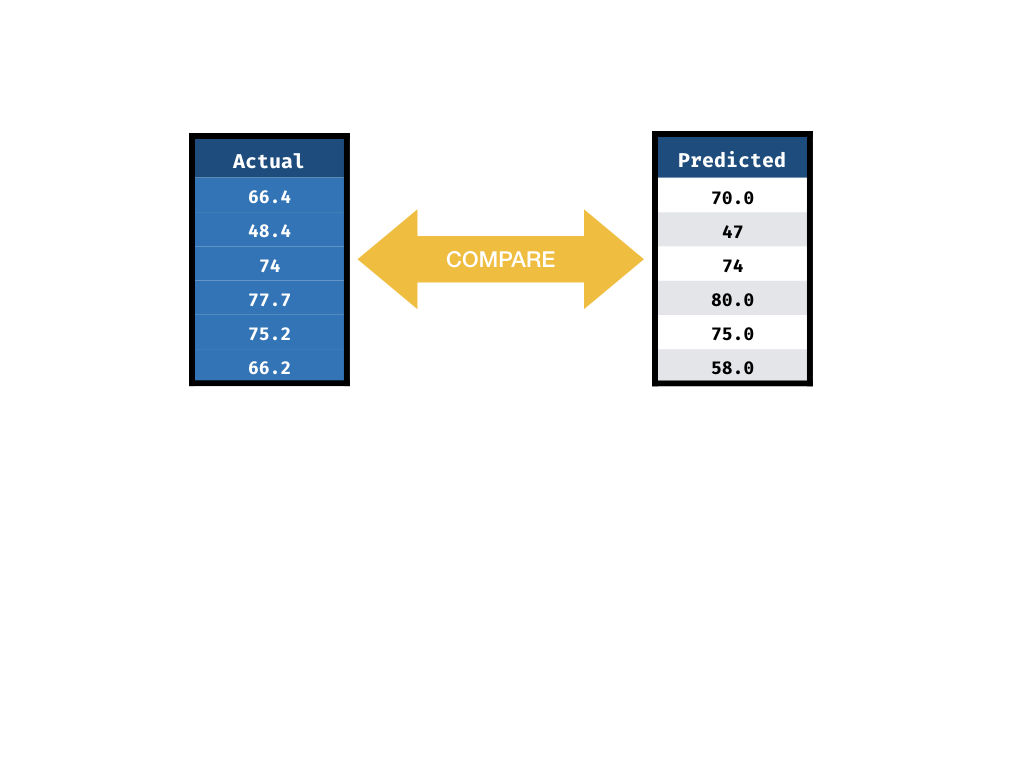
Measuring Performance
Mean Absolute Error
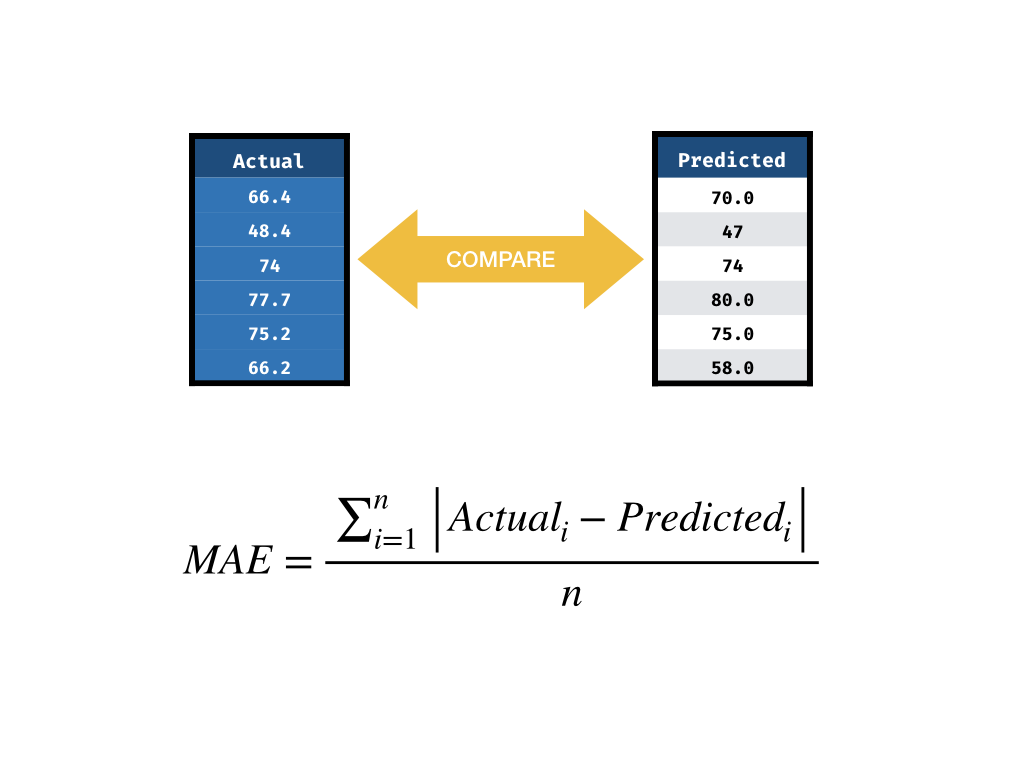
Ingredients for Performance Measurement
1) Actual life_expectancy values
2) Predicted life_expectancy values
3) A metric to compare 1) & 2)
1) Extract the actual values
cv_prep_lm <- cv_models_lm %>%
mutate(validate_actual = map(validate, ~.x$life_expectancy))
The predict() & map2() functions
predict(model, data)
map2(.x = model, .y = data, .f = ~predict(.x, .y))
2) Prepare the predicted values
cv_prep_lm <- cv_eval_lm %>%
mutate(validate_actual = map(validate, ~.x$life_expectancy),
validate_predicted = map2(model, validate, ~predict(.x, .y)))
3) Calculate MAE
library(Metrics) cv_eval_lm <- cv_prep_lm %>% mutate(validate_mae = map2_dbl(validate_actual, validate_predicted, ~mae(actual = .x, predicted = .y)))cv_eval_lm
# 5-fold cross-validation
# A tibble: 5 x 8
splits id train validate model validate_a. validate_p validate_mae
<S3: rsplit> Fold1 <tib. <tib. <S3. <dbl. <dbl. 1.47
<S3: rsplit> Fold2 <tib. <tib. <S3. <dbl. <dbl. 1.51
<S3: rsplit> Fold3 <tib. <tib. <S3. <dbl. <dbl. 1.44
<S3: rsplit> Fold4 <tib. <tib. <S3. <dbl. <dbl. 1.48
<S3: rsplit> Fold5 <tib. <tib. <S3. <dbl. <dbl. 1.68
Let's practice!
Machine Learning in the Tidyverse

